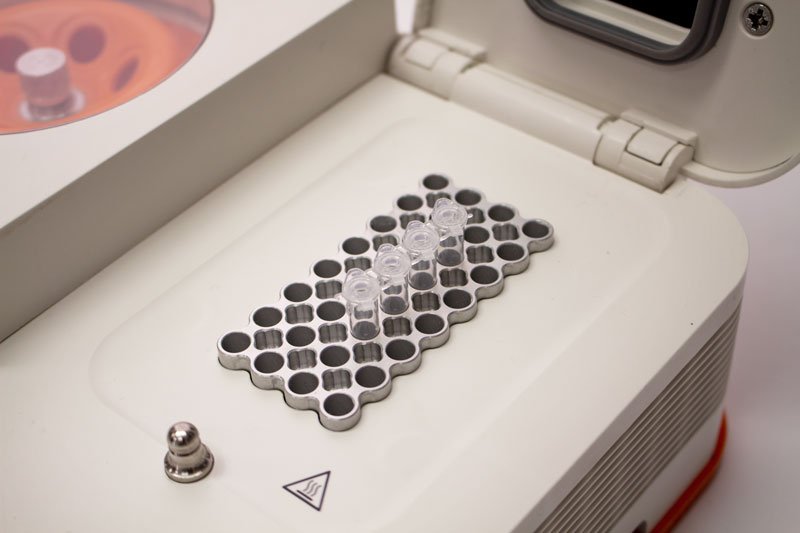
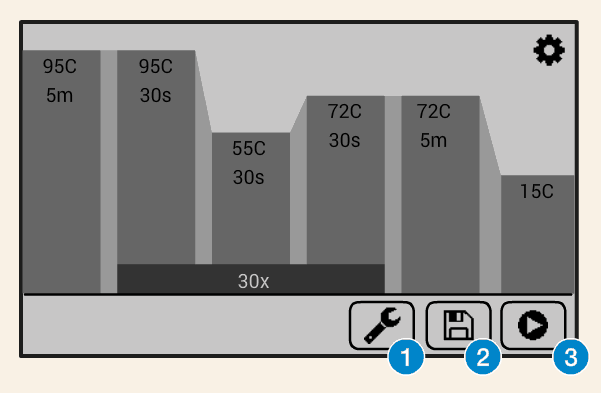
PCR (polymerase chain reaction) is a method used in molecular biology to make millions of physical copies of a specific DNA sequence, for example, a gene.
It has several key ingredients: a DNA template to copy, short DNA sequences called “primers”, and a master mix containing the rest of necessary molecules.
What is the template?
The DNA template is double-stranded genomic DNA from the sample.
What is a primer?
A primer is a short single strand of DNA that serves as a starting point for DNA synthesis of a new DNA strand. It is required for DNA replication because the enzymes that catalyze this process, DNA polymerases, can only add new nucleotides to an existing fragment of DNA.
How does PCR work?
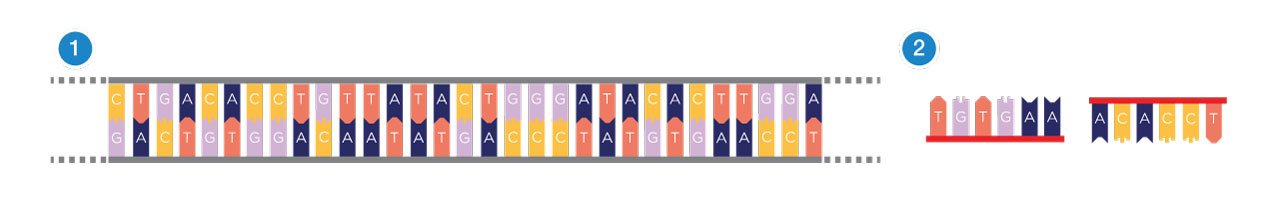
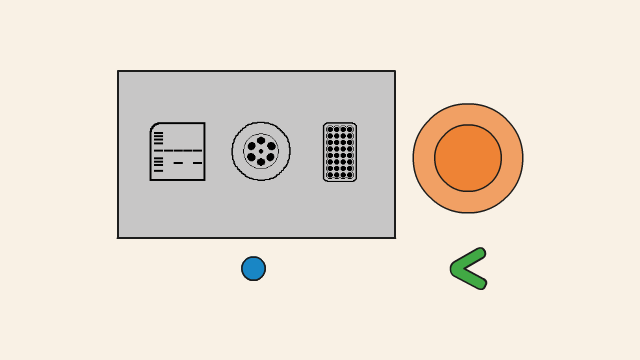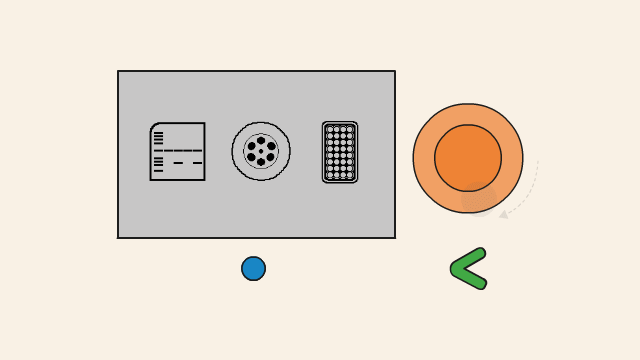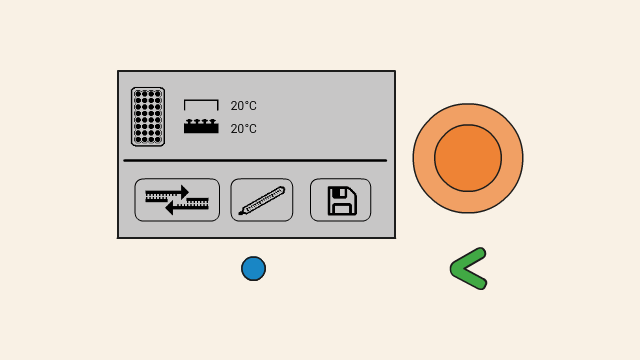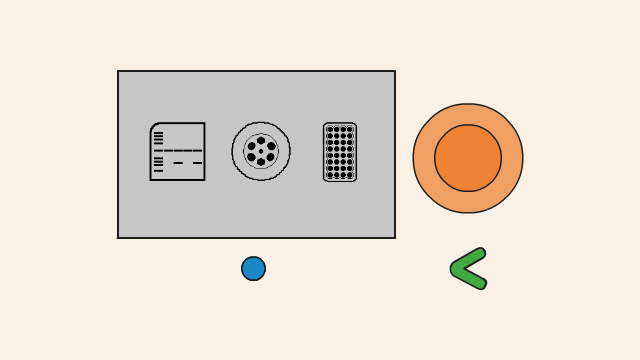
First of all, you need a DNA template (1), the physical DNA sample, which contains the sequence to be copied. It is a double stranded piece of DNA. You also need primers (2). These are short stretches of single-stranded DNA, which are designed to match the particular sequence of DNA you want to copy. Overall, you need four different primers that flank each side of the DNA you are trying to copy.
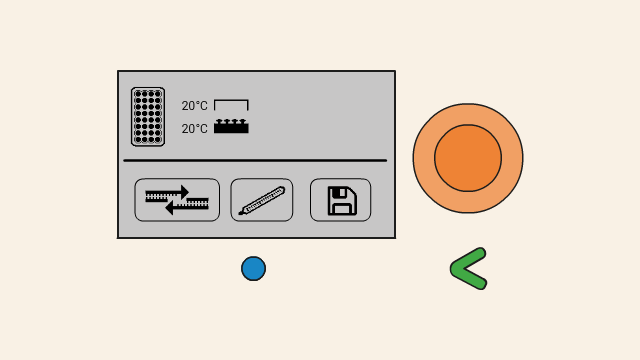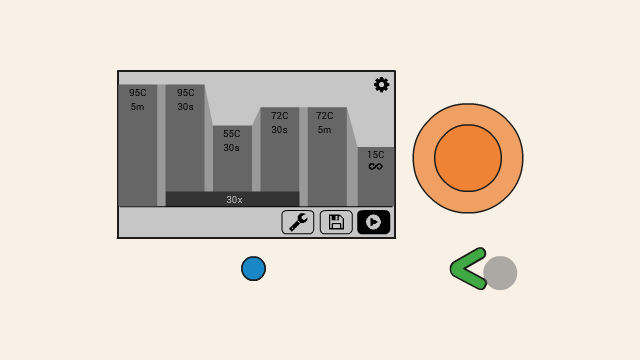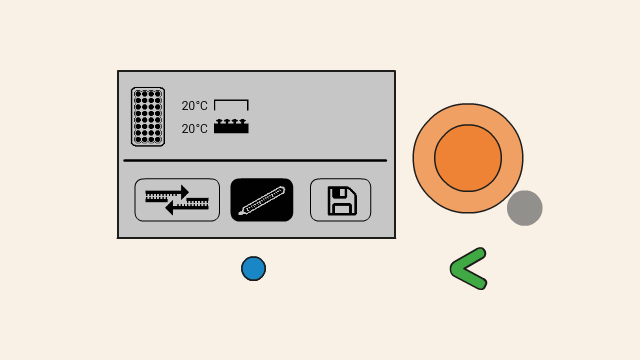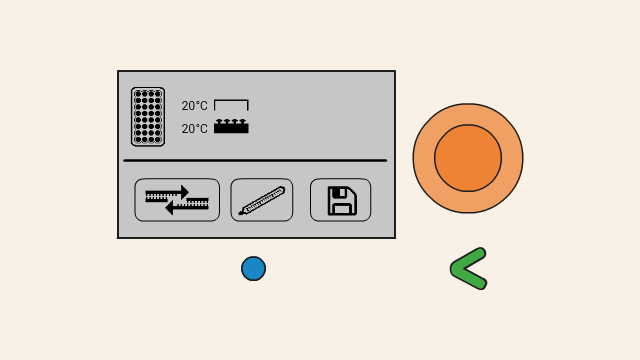
First Step: Denaturing
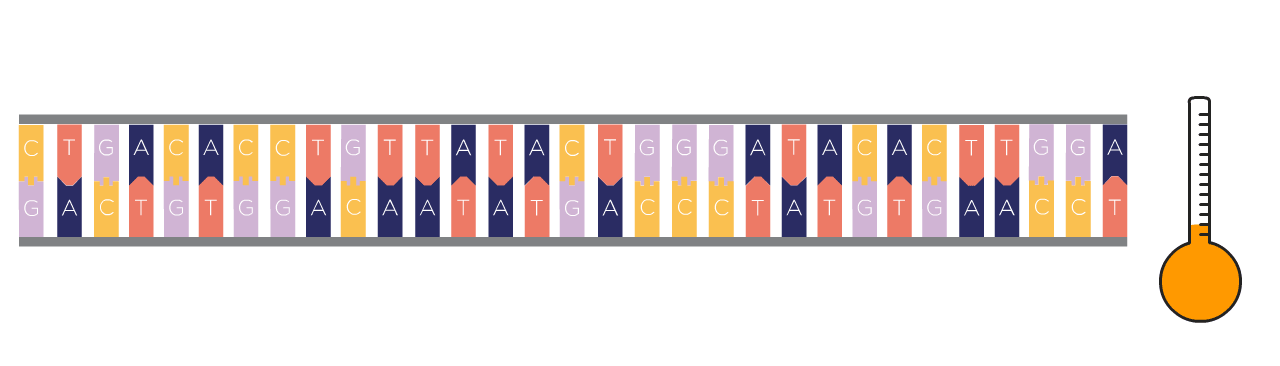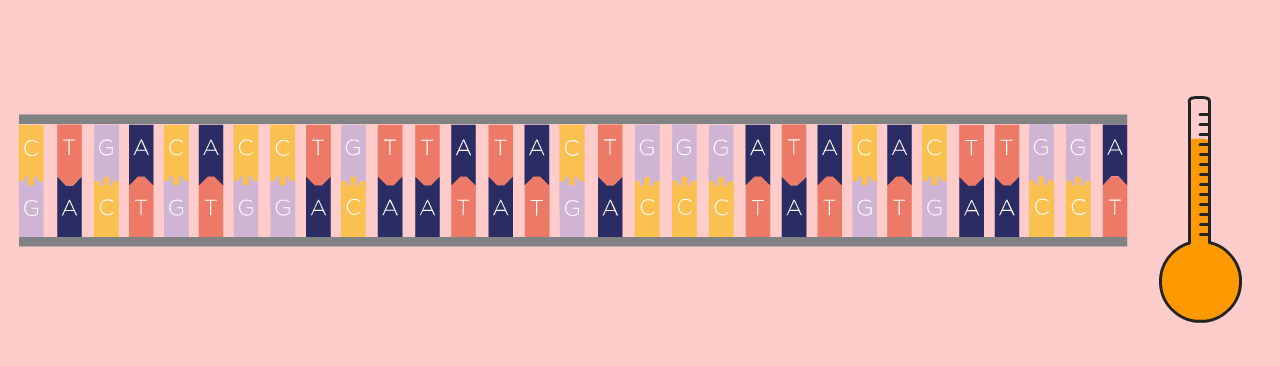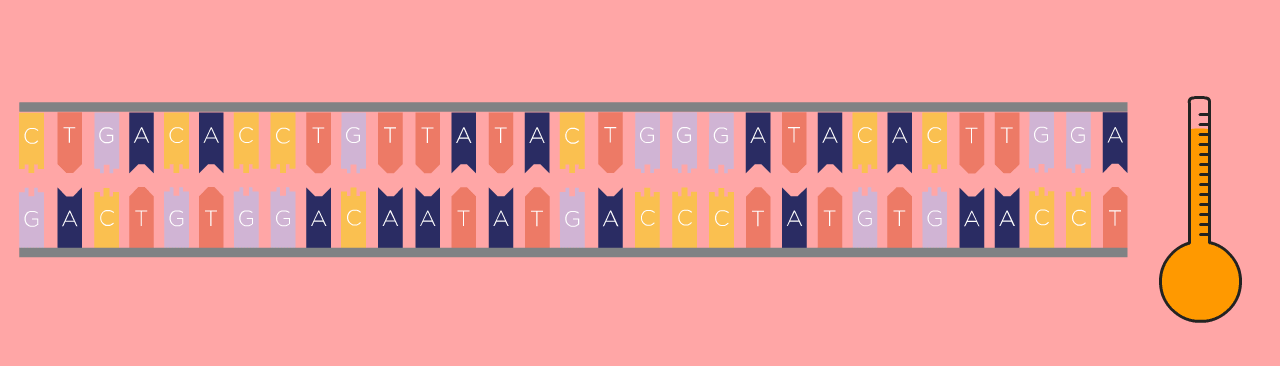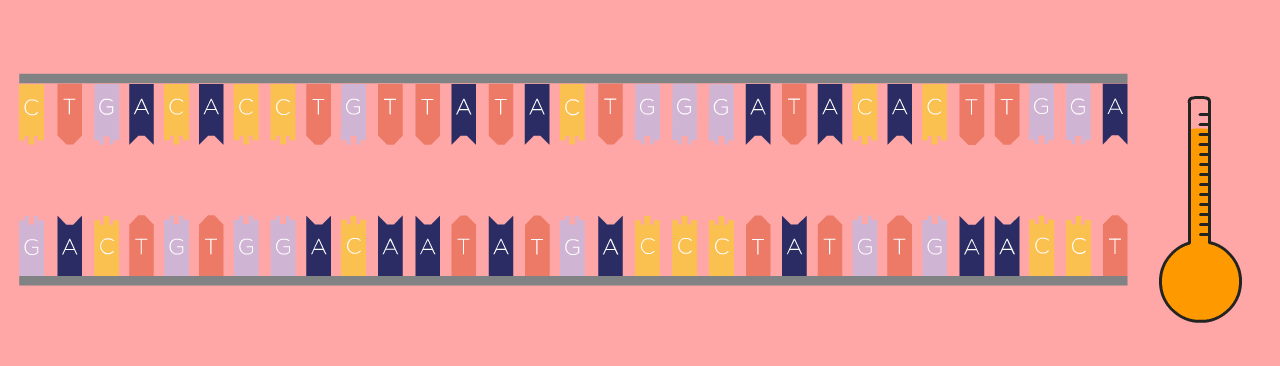
First, the DNA template is heated, which causes the double-stranded DNA to separate into two single strands.
Second Step: Annealing

Then the temperature is lowered to a temperature between 46-60 °C. The exact temperature depends on the experiment, and is known as the annealing temperature. At this point, the primers (2) will attach, or anneal, to their binding positions on the single strands of the template DNA.
Third Step: Extending
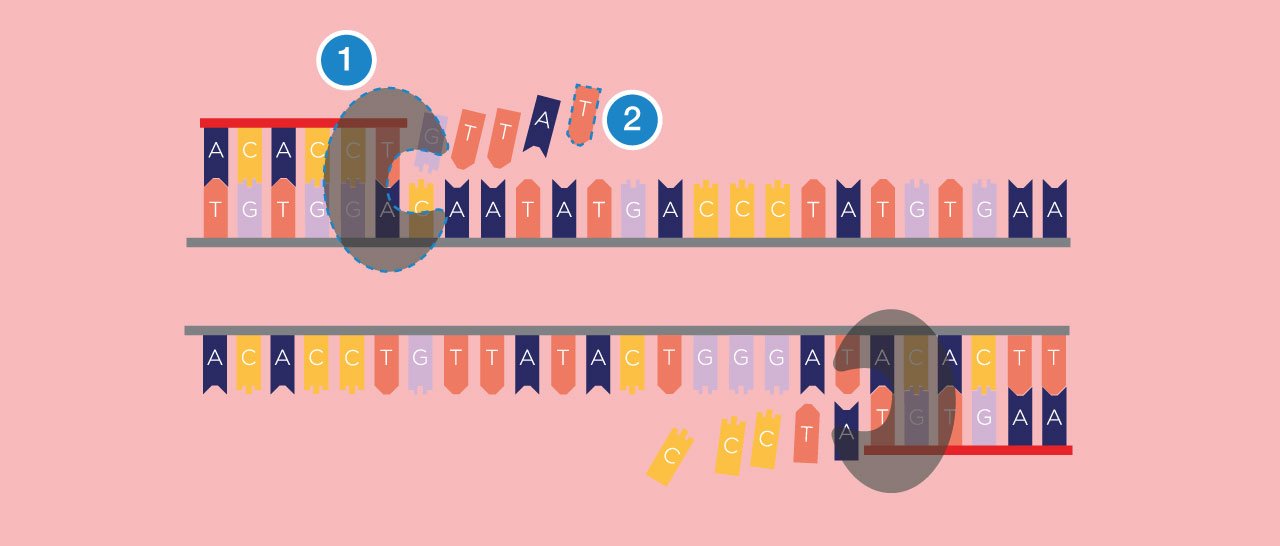
In the next step, the reaction moves to an optimal temperature for the polymerase enzyme (1), from which PCR takes its name, to start working. The polymerase enzyme builds DNA strands, and it will extend the DNA from the primer along the DNA template, creating a new DNA strand, which combines with the single-stranded template to form a double strand. The polymerase enzyme uses dNTPs (2), free DNA nucleotide bases as the building blocks for the new strand.
Repeated cycles for exponential growth
This process is repeated about 30-35 times. Each cycle doubles the copies of the DNA targeted by the primers.
Why PCR is so useful for analysis
Note that PCR will only work, if the template DNA contains the sequence you are targeting with the primers. This allows for the design of identification experiments. If some primers do not work, it is possible that the DNA sample did not contain the right DNA for the targeted sequence.